Bayesian Autoresearch for Causal Inference: When You Can’t Score the Thing You Care About
I’ve been exploring how the data scientist’s role shifts from building models to designing the systems that build them. I call this Agentic Data Science: you define objective functions, encode domain constraints in plain language, architect the experimentation loop. Then you step back.
This post is a concrete test of that idea. I built a system where a Claude agent autonomously iterates on Bayesian causal models. I specified everything in English: the objective, the constraints, the search strategy. Then I went to sleep.
By morning it had run 22 experiments. Nine crashed. The rest told a story about what happens when an AI agent optimizes a proxy for something it can never directly score.
Why causal inference is the hard test
Andrej Karpathy’s autoresearch showed this pattern works for ML: write training code, check if the loss dropped, iterate. The feedback loop is clean. Loss is a number. Lower is better. You can measure it on held-out data.
Causal inference breaks that loop.
When you estimate a treatment effect, the quantity you care about is a counterfactual. You never observe both potential outcomes for the same person. There is no test set for causal effects. The Average Treatment Effect (ATE) is not a prediction error you can compute. It’s a belief about a world you can’t see.
So I needed a proxy. Since you can’t score the treatment effect directly, you have to design a heuristic: a set of metrics and rules that you believe will push the model in the right direction. This is a design choice, not something you hand off to the AI. Other practitioners might choose different proxies or different thresholds. The specification I gave the agent, in plain English:
- Use ELPD (expected log pointwise predictive density via PSIS-LOO) to score predictive accuracy
- Gate on convergence: R-hat < 1.01, ESS > 400, divergence rate < 0.1%
- Keep a new model only if ELPD improves by more than 2 standard errors
- When two models tie on ELPD, prefer the one with a narrower credible interval on the treatment effect
The bet behind this heuristic: a model that captures the data-generating process well will also give good causal estimates. It’s a reasonable bet, one that Bayesian statisticians have been making for decades, but it’s not guaranteed. The question I wanted to answer is different: given this specification, can an LLM take it and iterate on its own?
What happened: 22 experiments on IHDP
The test bed is IHDP (Infant Health and Development Program): 747 observations, 25 confounders (6 continuous, 19 binary), binary treatment (home visits by specialist doctors), continuous outcome (infant cognitive test scores). It’s semi-synthetic, meaning both potential outcomes exist and the true ATE (~4.0) is known. In a real problem you wouldn’t have that check.
The agent ran 22 experiments overnight.
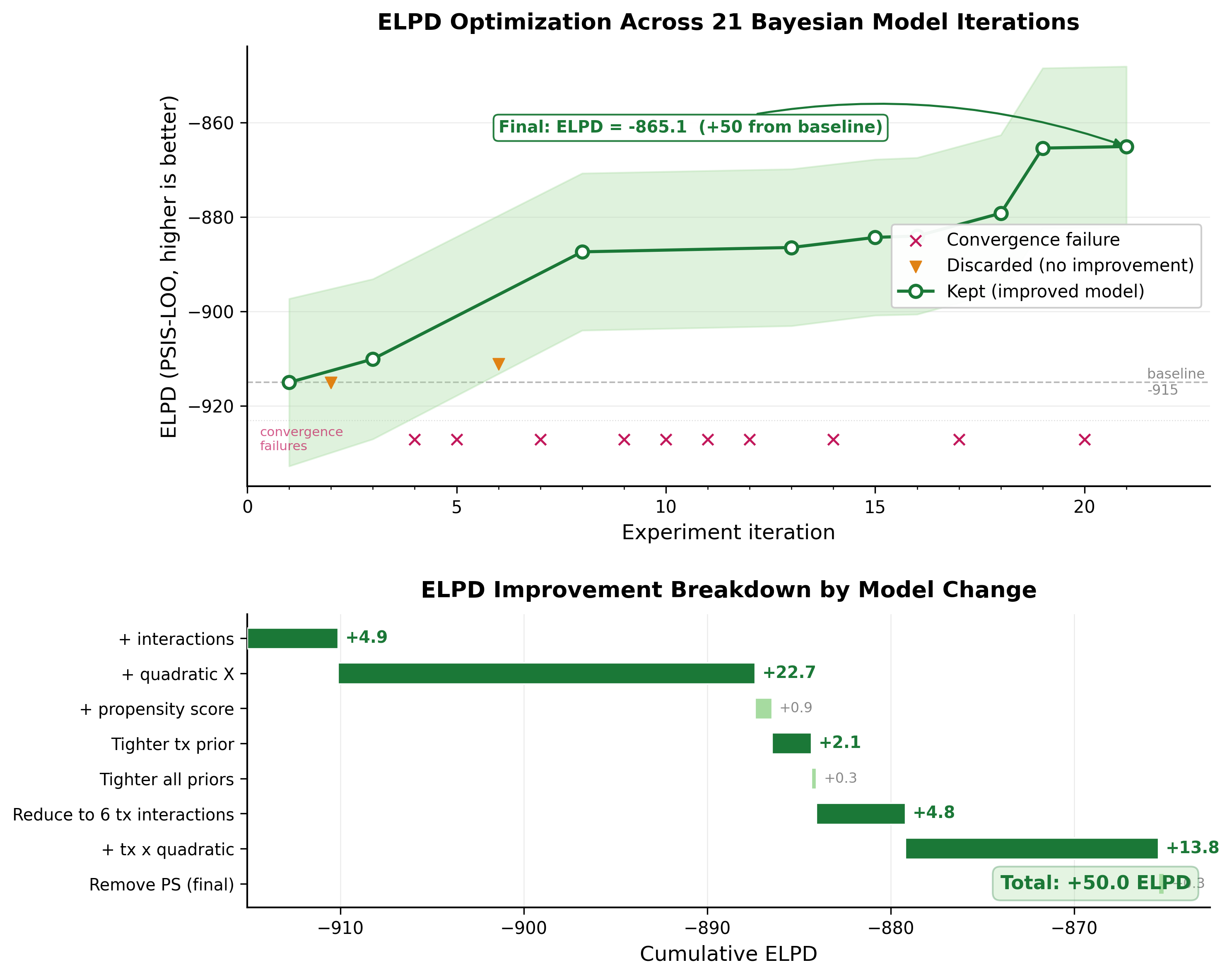
Two changes dominated: adding quadratic confounders (+22.7 ELPD) and treatment-by-quadratic interactions (+13.8 ELPD). The smaller steps (tightening priors, adding propensity scores) contributed less than 2 ELPD points each. Total improvement: 50 points from baseline.
But ELPD is the proxy. The real question is what happened to the causal estimate.
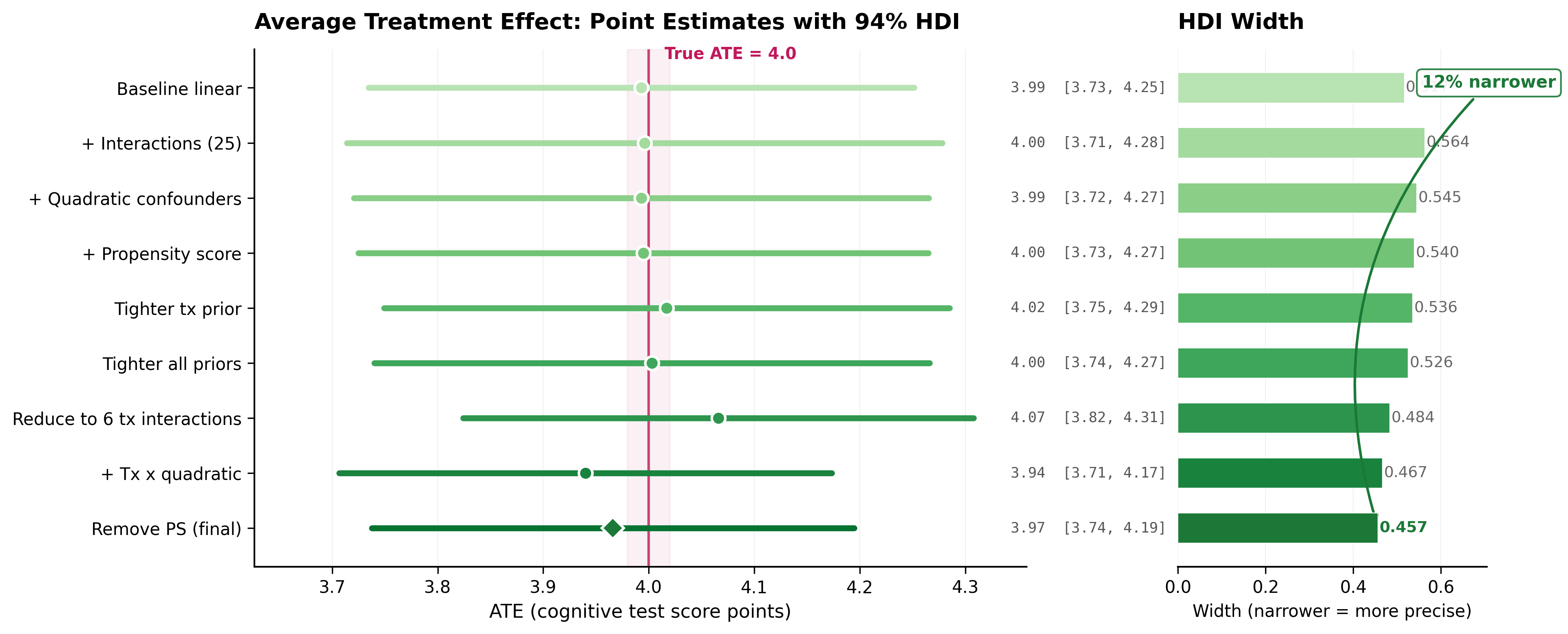
Every model’s ATE estimate sits close to the true value of 4.0, and every credible interval contains it. But the intervals get tighter as the models improve, from 0.517 down to 0.457 (12% narrower).
The path wasn’t monotonic. Adding 25 treatment-confounder interaction terms (experiment 3) improved predictions but made the credible interval wider, from 0.517 to 0.564. More parameters meant more uncertainty in the treatment effect. The real precision gain came when the agent restricted interactions to only the 6 continuous confounders, dropping 19 binary ones that don’t carry enough variation in 747 rows to support interaction terms.
The agent reasoned through failures too. Treatment-quadratic interactions crashed at experiment 9 with 53 parameters (R-hat > 1.01). Instead of abandoning the idea, it simplified the base model first, then re-introduced the same interactions on the simpler foundation. It worked. The agent connected “too many parameters caused convergence failure” to “simplify before adding complexity.” This is the same pattern that makes LLM coding agents effective: try something, hit a wall, read the error, form a new hypothesis, try again. Except here the errors are convergence diagnostics instead of stack traces, and the hypotheses are statistical models instead of code patches. The agent remembers what it tried, why it failed, and uses that to make educated guesses about what to try next.
9 of 22 experiments crashed. The 40% failure rate is the system working: the agent proposes, the evaluation harness vetoes.
How the system works
So how does the system actually work?
The system’s configuration lives in two prompt files, both written in English. This is where the data scientist focuses.
1. PROGRAM.md defines the experimentation procedure: the objective function, the keep/discard rule, and a search heuristic (“build complexity incrementally; if stuck, simplify or combine elements from previous near-misses”). 2. PROBLEM.md encodes problem-specific domain knowledge: the causal question, the variable structure, and business logic that only a domain expert would know, like “column x4 has data leakage, do not use it” or “there is no causal path between X1 and Y.”
General statistical knowledge (when to use non-centered parameterizations, how interactions affect CATE estimation) does not belong in these two files. It belongs in a domain-specific skill given to the execution agents (see the pymc-modeling skill below). The separation matters: change PROBLEM.md and the same system works on a different dataset. Change PROGRAM.md and you get a different search strategy. Swap the skill and you get a different modeling paradigm entirely.
Bayesian modeling fits this pattern particularly well. The data scientist specifies knowledge in natural language, and the LLM takes care of translating these constraints into mathematical priors written directly in code.
Three components execute the loop:
The orchestrator is a Claude Code session running an infinite loop. It reads PROGRAM.md and past experiment results, reasons about what to try next, and spawns a coding sub-agent with a specific hypothesis.
The coding sub-agent gets a fresh context each iteration. It reads PROBLEM.md, the current model, and the experiment history, then edits a single file (model.py). It loads a pymc-modeling skill that gives it statistical modeling expertise: when non-centered parameterizations help, how priors interact with sample size, what diagnostic values mean. The agent isn’t a generic coder. It’s a coder with a statistics textbook in working memory, reading a problem brief written by the domain expert.
The evaluation harness is fixed Python code I wrote once and never touched. Data splitting, MCMC sampling with nutpie (5-minute timeout), convergence diagnostics, ELPD computation, ATE estimation. If the model doesn’t converge, crash and revert. If it converges but isn’t better, discard.
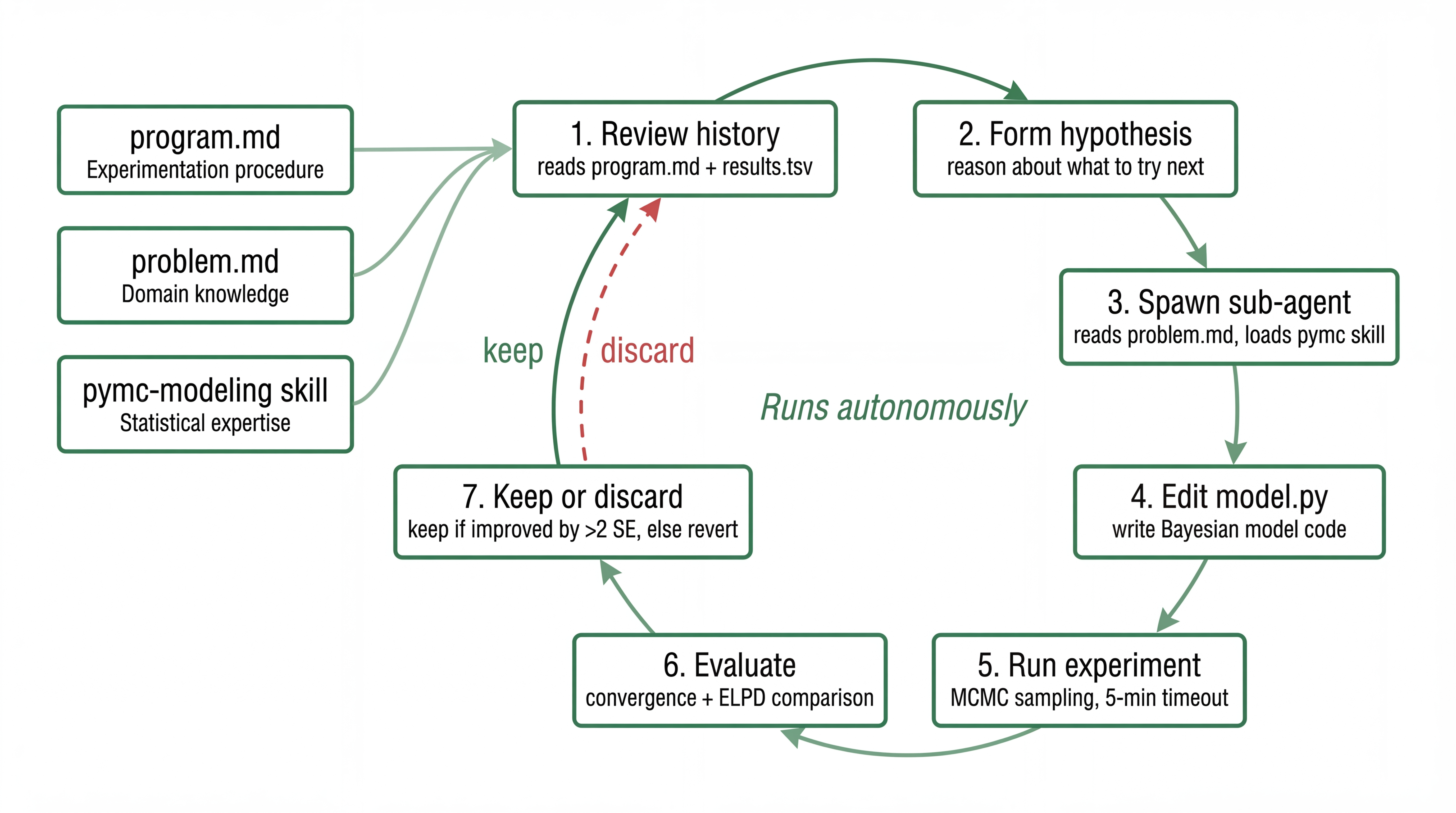
Note that the verification loop is encoded naturally here. Convergence diagnostics tell the agent when something is wrong without a human looking at trace plots. Credible intervals quantify uncertainty on the causal estimate. The math itself is a feedback channel.
Did the bet pay off?
Final ATE: 3.97, within 1.3% of the ground truth. The 94% credible interval [3.74, 4.19] comfortably contains the true value. Zero divergences, R-hat at 1.00, ESS above 700.
\[ \mu_i = \alpha + \beta_t \cdot t_i + \mathbf{X}_i \boldsymbol{\beta}_x + t_i \cdot \mathbf{X}^{cont}_i \boldsymbol{\beta}_{tx} + \mathbf{X}^{2}_{cont,i} \boldsymbol{\beta}_{sq} + t_i \cdot \mathbf{X}^{2}_{cont,i} \boldsymbol{\beta}_{tx,sq} \]
| Parameter | Dim | Prior | Role |
|---|---|---|---|
| \(\alpha\) | 1 | \(N(0, 3)\) | Intercept |
| \(\beta_t\) | 1 | \(N(0, 2)\) | Treatment effect (ATE base term) |
| \(\boldsymbol{\beta}_x\) | 25 | \(N(0, 1)\) | Linear confounder adjustment (all 25) |
| \(\boldsymbol{\beta}_{tx}\) | 6 | \(N(0, 0.7)\) | Treatment \(\times\) continuous confounders |
| \(\boldsymbol{\beta}_{sq}\) | 6 | \(N(0, 0.5)\) | Quadratic confounders \((x_1\text{-}x_6)^2\) |
| \(\boldsymbol{\beta}_{tx,sq}\) | 6 | \(N(0, 0.3)\) | Treatment \(\times\) quadratic confounders |
| \(\sigma\) | 1 | \(\text{HalfNormal}(2)\) | Observation noise |
All 25 confounders enter main effects, but only the 6 continuous ones get treatment interactions and quadratic terms. Priors get progressively tighter from main effects (1.0) to linear interactions (0.7) to quadratic interactions (0.3), encoding that higher-order terms should be smaller.
I reviewed this after the fact and would have probably made similar choices, though it would have taken me two days of manual iteration rather than one night. The agent also tried ideas I might not have bothered with: horseshoe priors on interactions (1,368 divergences), cubic confounders, pairwise confounder interactions, Laplace priors. Some of that exploration was wasteful. But it’s also how the agent discovered that quadratic interactions work, just on a simpler foundation than it first tried.
I also ran the same system on a second dataset: Twins (twin birth mortality, ~5000 observations, binary outcome). Same PROGRAM.md, different PROBLEM.md. 32 experiments, 10 kept, 9 crashed. ELPD went from 1579 to 3139 (99% improvement), ATE HDI width from 0.024 to 0.011 (53% narrower). The agent discovered that outcome variance depends on gestational age (heteroscedastic modeling), a sophisticated move. Different dataset, different model family, same pattern.
The honest caveat: both experiments are on semi-synthetic benchmarks. IHDP and Twins are well-behaved. The confounding is strong but not pathological. And the baseline models are already quite strong on these datasets, coming close to the true ATE even before the agent starts iterating. The improvements are real but incremental on problems where a simple linear model already does most of the work. Real-world data with unmeasured confounders or positivity violations would be a harder test.
Principles of agentic data science
These emerged from building the system. They aren’t specific to Bayesian causal inference but also apply to traditional predictive modeling and ML.
Objective as specification. The human defines what “good” means: metrics, constraints, comparison rules. The agent optimizes. I chose ELPD as primary, convergence as gate, HDI width as tiebreaker. That choice is itself a domain judgment, and it’s the thing that matters most.
Domain knowledge as natural language. Causal structure, prior beliefs, business constraints, all written in English. The LLM translates them to probabilistic code and priors. That translation is what makes the whole thing work. LLMs read English and write PyMC models. They read convergence diagnostics and form new hypotheses.
Technical expertise as loadable skills. The pymc-modeling skill gives the agent statistical competence, reusable across problems. The agent isn’t general-purpose. It’s equipped with best practices and opinionated patterns.
Experimentation procedure as a loop. The search strategy is written in words (“start simple, add complexity, if stuck try simplifying”) that the agent interprets and adapts as experiments succeed or fail. Each sub-agent starts with a fresh context containing only what worked and what didn’t, keeping the working memory lean while still accumulating knowledge across iterations.
Guardrails and verification. I defined what the agent must not do (convergence gates, time budgets, significance tests) in instructions provided to the agent. Then I checked the final model for plausibility, not every intermediate experiment. The agent explores freely within a constrained space. If it had produced something implausible that happened to score well, some of these constraints would have caught it. This prototype relies on the agent’s instruction-following ability; production-grade implementations can enforce the same constraints with hooks and callbacks.
What this is and what it isn’t
This is a proof of concept focusing on one slice of the causal data science workflow: model iteration. Real data science involves data cleaning, feature engineering, sensitivity analysis, outcome definition, stakeholder alignment, deployment monitoring. The Agentic Data Science article discusses the full picture. Many of those steps matter more than the model specification itself.
One natural objection: English is ambiguous. Imprecise specifications lead to wrong models. True, but this is an engineering problem. Production systems can add verification loops where the agent flags ambiguity and asks for clarification. And Bayesian modeling helps: the math surfaces problems. A posterior that’s too wide, a model that won’t converge, a prior that conflicts with the data. These are signals the agent can act on, or escalate to the human.
The code is on GitHub. It ships with four causal inference problems, but the architecture is general. If you have a dataset and a causal question, swap the PROBLEM.md and point it at your data. I’m curious what happens on problems where nobody knows the right answer.